The University of the Philippines chemists have developed a new artificial intelligence tool. It speeds up the discovery of new antibacterial treatments with the aim of preventing global antimicrobial resistance.
Antimicrobial resistance makes traditional antibiotics less effective. This creates a need for new medical solutions. Researchers are now looking at antimicrobial peptides. These are small molecules capable of killing bacteria.
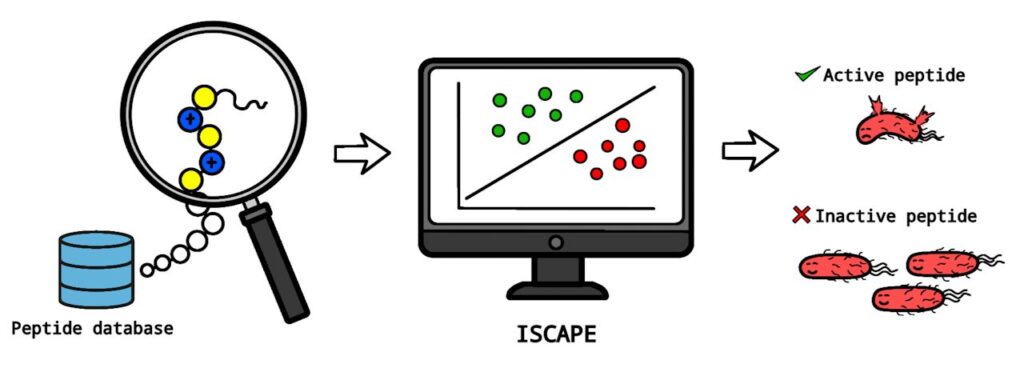
Remmer Salas, Dr. Portia Mahal Sabido, and Dr. Ricky Nellas of the Institute of Chemistry developed the tool. They named it ISCAPE. The name stands for Interpretable Support Vector Classifier of Antibacterial Activity of Peptides against Escherichia coli.
The tool predicts if a peptide can kill or stop the growth of E. coli. Users only need to provide a Simplified Molecular-Input Line-Entry System string to get results.
“Traditionally, discovering antibacterial peptides means synthesizing many candidates and testing them one by one. This process is time-consuming,” Salas said. “We used AI to learn from existing data and identify patterns that distinguish active peptides from inactive ones.”
The tool differs from other AI models because it shows exactly which molecular features make a peptide effective. This insight helps scientists save time and resources. It lowers the need for trial-and-error laboratory experiments.
“ISCAPE helps address antimicrobial resistance by accelerating early-stage screening through data-driven peptide design,” Salas explained. “It doesn’t replace laboratory experiments, but it makes discovery more efficient and helps researchers focus on the most promising candidates.”
The model is flexible. It can be adapted to target other types of bacteria. However, the system requires retraining with high-quality datasets for specific bacterial strains. Researchers could also use the ISCAPE approach to predict other types of bioactive peptides.
“Applying ISCAPE to other biological targets requires well-curated datasets with experimentally validated activity,” Salas added. “The model must then be retrained using the molecular features we identified as optimal for peptides.”
The research team aims to help to the global effort against antimicrobial resistance. They want to help scientists design better peptides with greater efficiency.
This tool is available to the public. Scientists can access the web server through Hugging Face Spaces. The code and training data are also available on GitHub.
The team published their findings in the Journal of Molecular Graphics and Modelling. The study is titled “Interpretable support vector classifier for reliable prediction of antibacterial activity of modified peptides against Escherichia coli.”
This international journal focuses on the use of computers in molecular chemistry. It covers molecular structure, function, interaction, and design.